Plot classifier metrics as a function of thresholds.
ThresholdPlot( frame, xvar, truthVar, title, ..., metrics = c("sensitivity", "specificity"), truth_target = TRUE, points_to_plot = NULL, monochrome = TRUE, palette = "Dark2", linecolor = "black" )
Arguments
| frame | data frame to get values from |
|---|---|
| xvar | column of scores |
| truthVar | column of true outcomes |
| title | title to place on plot |
| ... | no unnamed argument, added to force named binding of later arguments. |
| metrics | metrics to be computed. See Details for the list of allowed metrics |
| truth_target | truth value considered to be positive. |
| points_to_plot | how many data points to use for plotting. Defaults to NULL (all data) |
| monochrome | logical: if TRUE, all subgraphs plotted in same color |
| palette | character: if monochrome==FALSE, name of brewer color palette (can be NULL) |
| linecolor | character: if monochrome==TRUE, name of line color |
Details
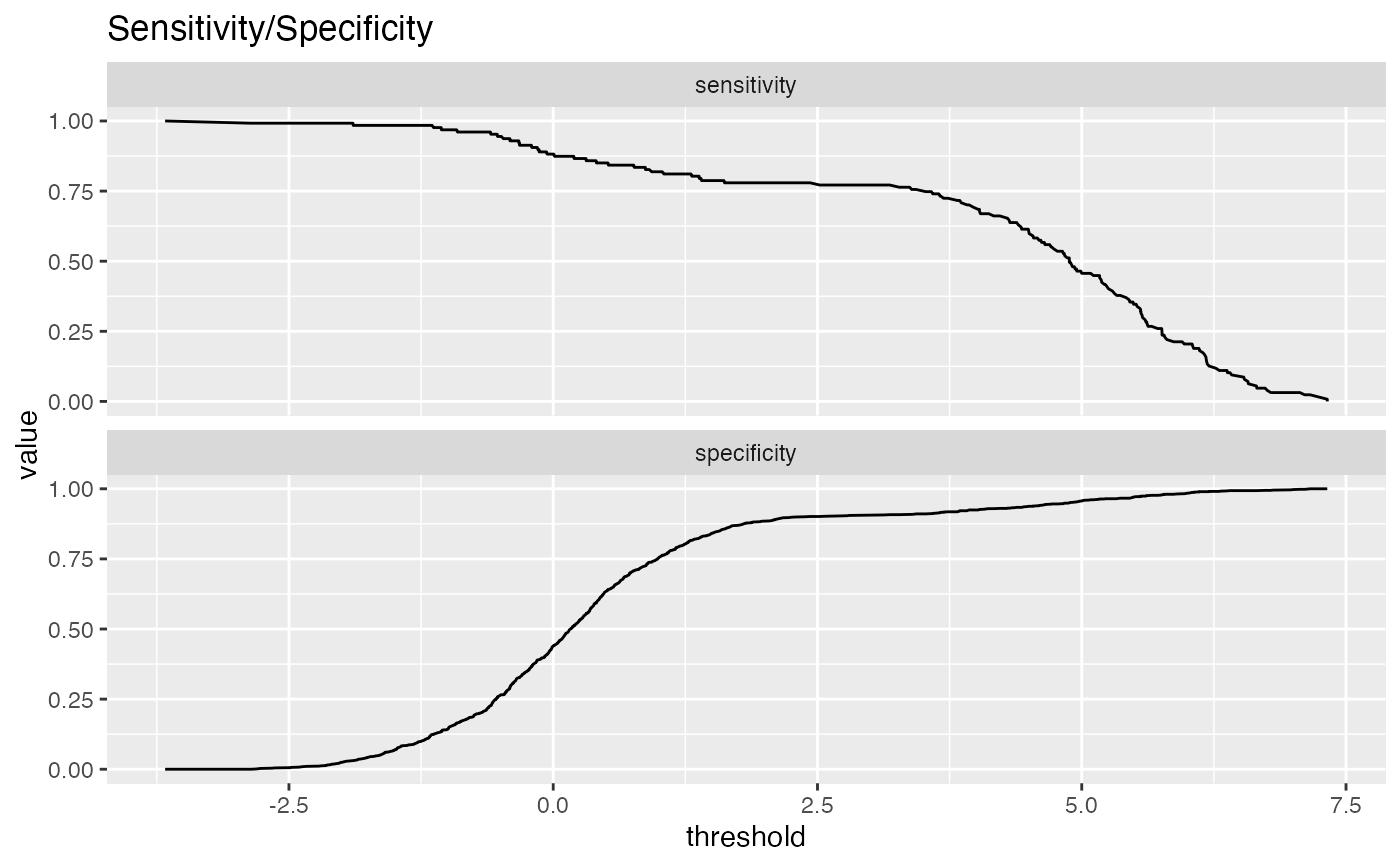
By default, ThresholdPlot plots sensitivity and specificity of a
a classifier as a function of the decision threshold.
Plotting sensitivity-specificity (or other metrics) as a function of classifier score helps
identify a score threshold that achieves an acceptable tradeoff among desirable
properties.
ThresholdPlot can plot a number of metrics. Some of the metrics are redundant,
in keeping with the customary terminology of various analysis communities.
sensitivity: fraction of true positives that were predicted to be true (also known as the true positive rate)
specificity: fraction of true negatives to all negatives (or 1 - false_positive_rate)
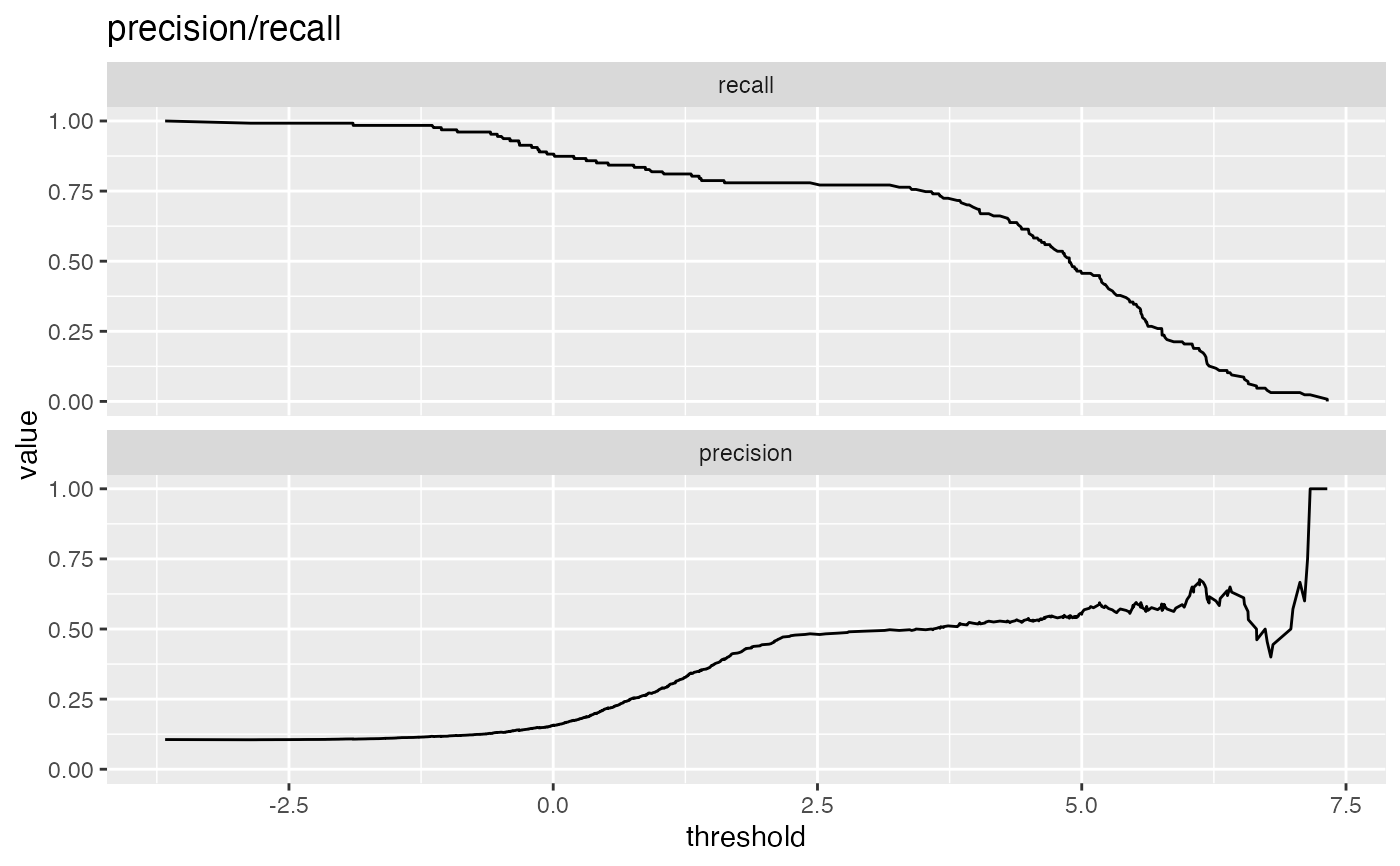
precision: fraction of predicted positives that are true positives
recall: same as sensitivity or true positive rate
accuracy: fraction of items correctly decided
false_positive_rate: fraction of negatives predicted to be true over all negatives
true_positive_rate: fraction of positives predicted to be true over all positives
false_negative_rate: fraction of positives predicted to be all false over all positives
true_negative_rate: fraction negatives predicted to be false over all negatives
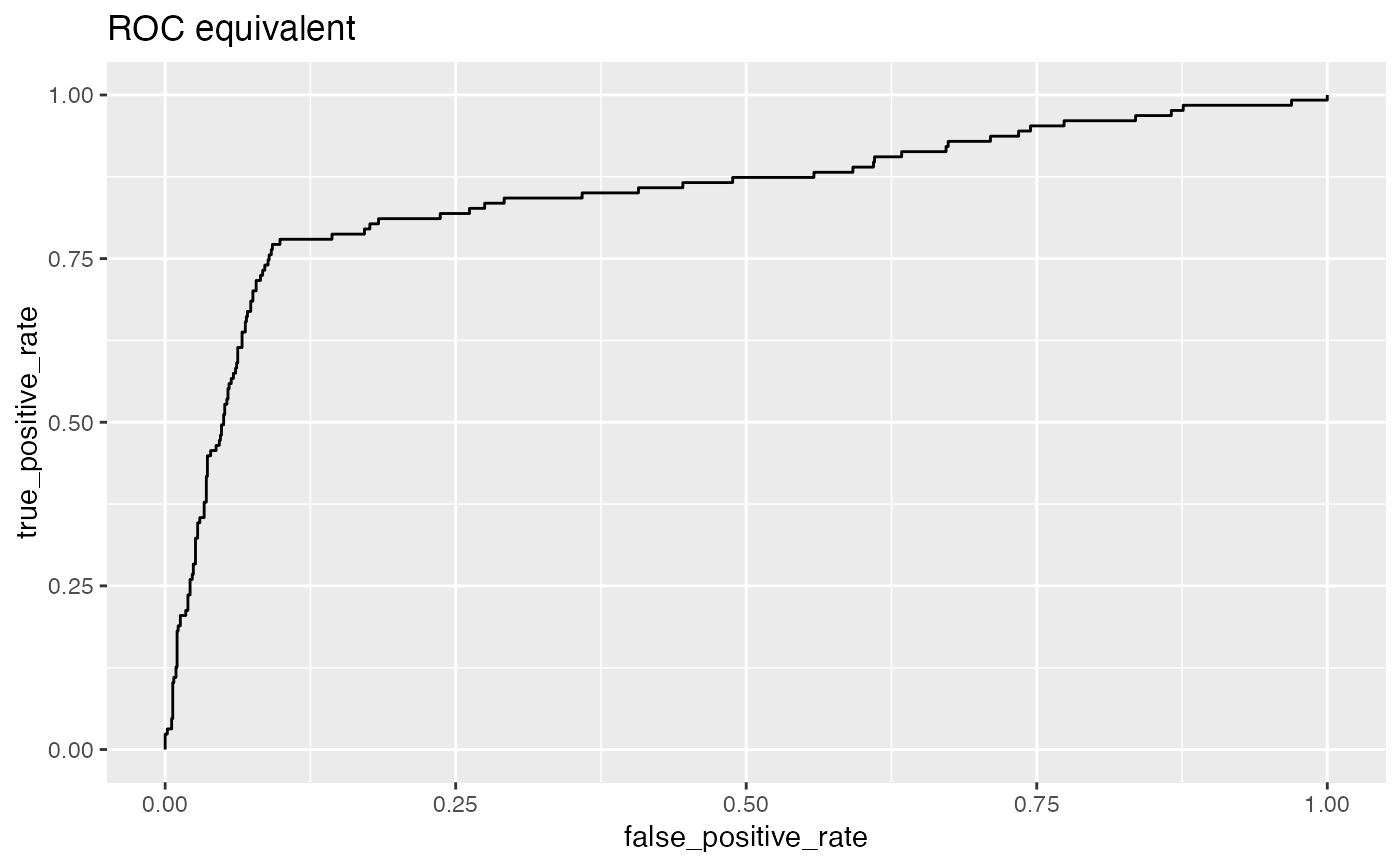
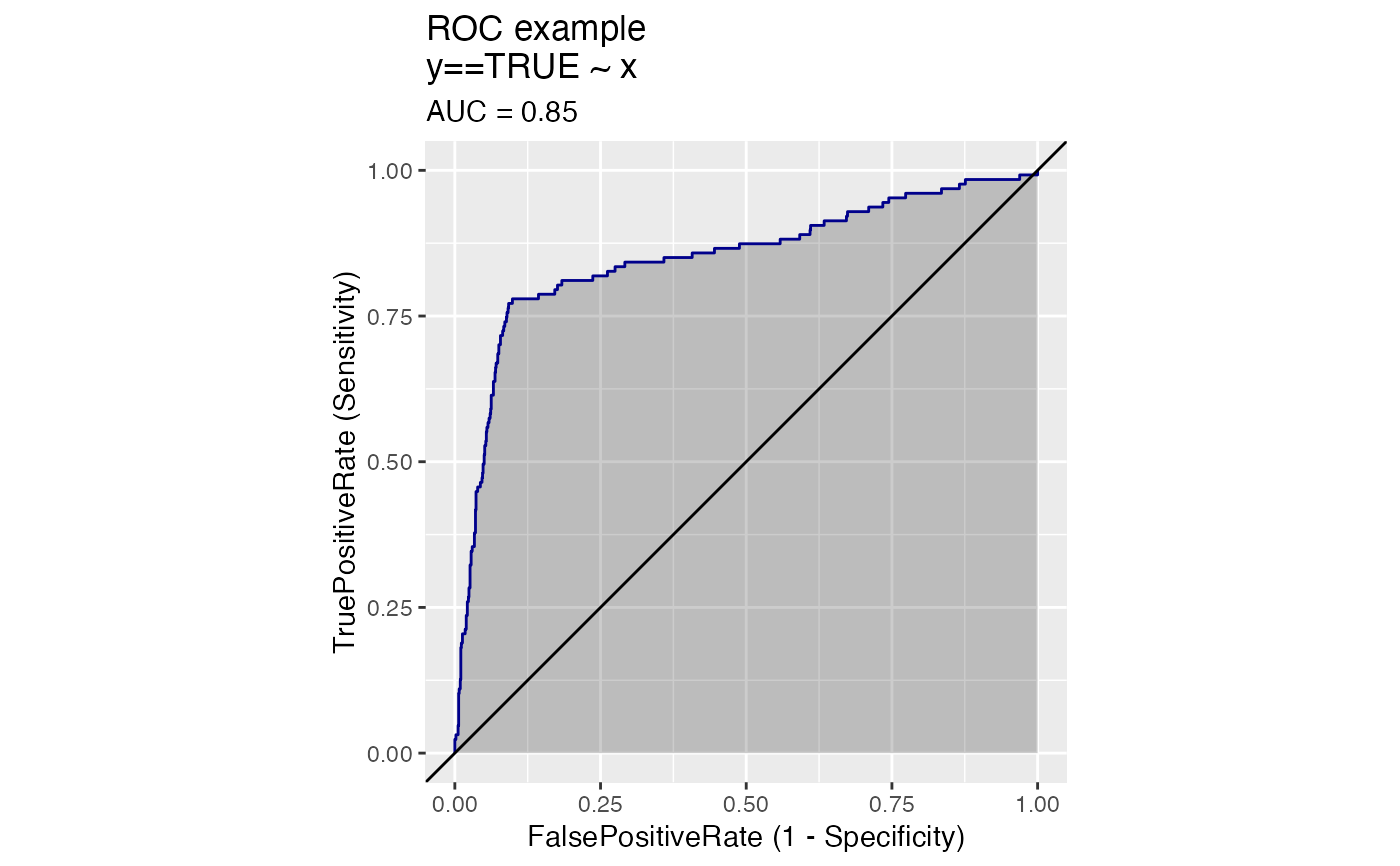
For example, plotting sensitivity/false_positive_rate as functions of threshold will "unroll" an ROC Plot.
ThresholdPlot can also plot distribution diagnostics about the scores:
fraction: the fraction of datums that scored greater than a given threshold
cdf: CDF or
1 - fraction; the fraction of datums that scored less than a given threshold
Plots are in a single column, in the order specified by metrics.
points_to_plot specifies the approximate number of datums used to
create the plots as an absolute count; for example setting points_to_plot = 200 uses
approximately 200 points, rather than the entire data set. This can be useful when
visualizing very large data sets.
See also
Examples
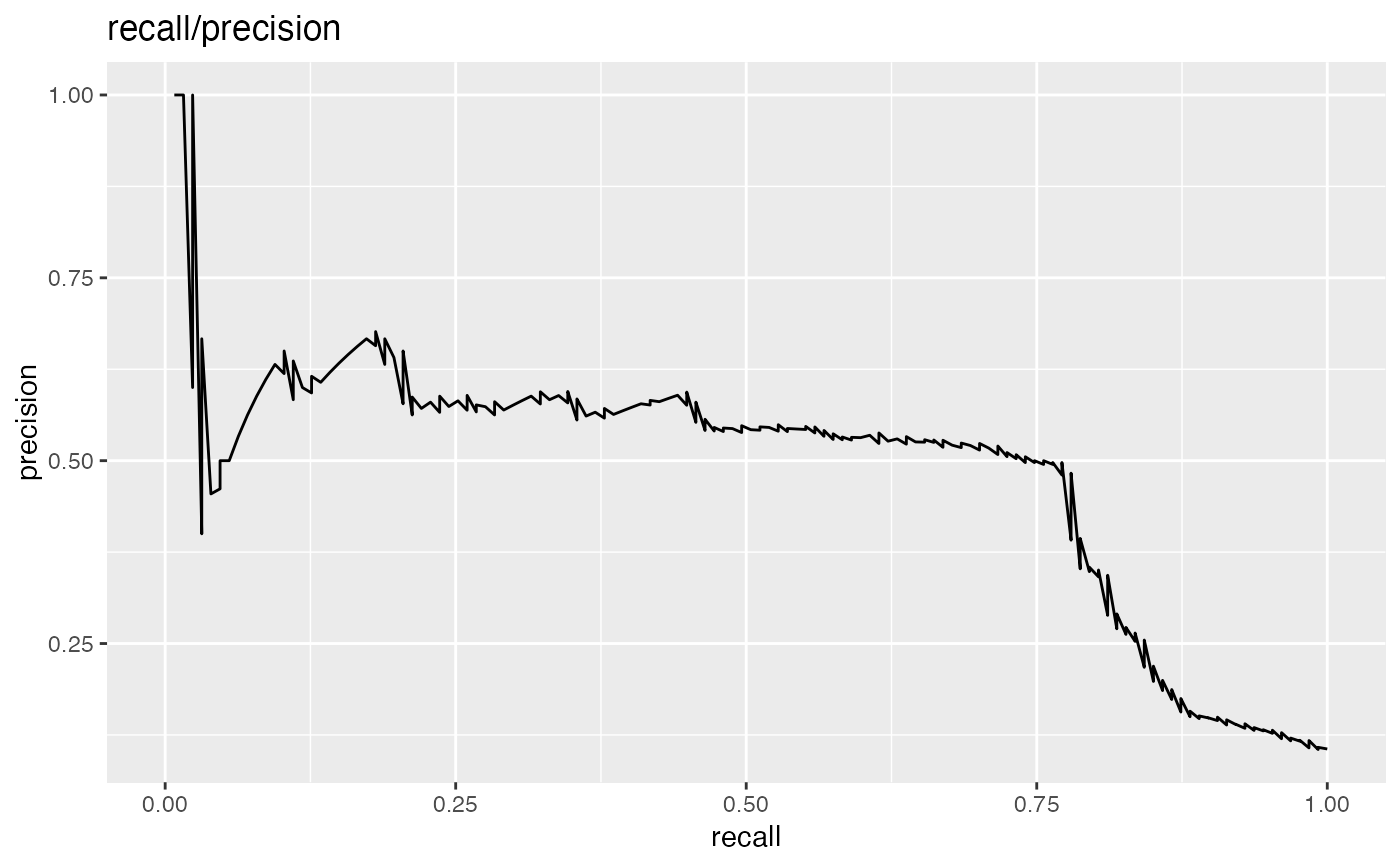
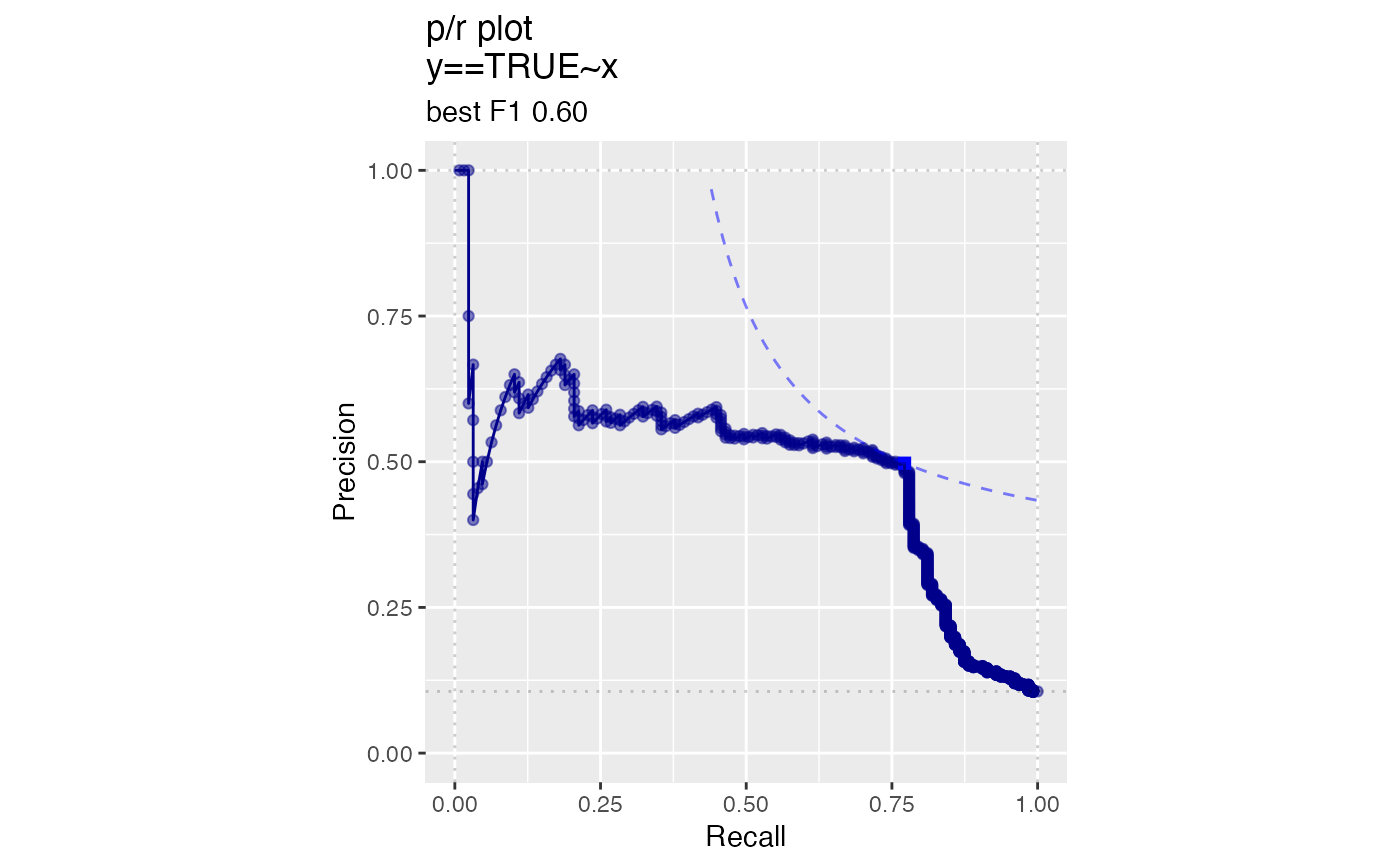
# data with two different regimes of behavior d <- rbind( data.frame( x = rnorm(1000), y = sample(c(TRUE, FALSE), prob = c(0.02, 0.98), size = 1000, replace = TRUE)), data.frame( x = rnorm(200) + 5, y = sample(c(TRUE, FALSE), size = 200, replace = TRUE)) ) # Sensitivity/Specificity examples ThresholdPlot(d, 'x', 'y', title = 'Sensitivity/Specificity', metrics = c('sensitivity', 'specificity'), truth_target = TRUE)MetricPairPlot(d, 'x', 'y', x_metric = 'false_positive_rate', y_metric = 'true_positive_rate', truth_target = TRUE, title = 'ROC equivalent')# Precision/Recall examples ThresholdPlot(d, 'x', 'y', title = 'precision/recall', metrics = c('recall', 'precision'), truth_target = TRUE)#> Warning: Removed 1 row(s) containing missing values (geom_path).MetricPairPlot(d, 'x', 'y', x_metric = 'recall', y_metric = 'precision', title = 'recall/precision', truth_target = TRUE)#> Warning: Removed 2 row(s) containing missing values (geom_path).